Genomic dissection of non-genetic cellular heterogeneity
One of the biggest challenges in biology is to understand how apparently identical cells respond differently to the same stimulus. In some cases, this differential behaviour can be explained by alterations of their genetic material, however in other cases identical (clonal) cells can also display phenotypically heterogeneous responses. We use state of the art genomic technologies to study the regulatory mechanism leading to the appearance of divergent gene expression programs in clonal population
In the last decades, the biomedical field has suffered a revolution thanks to the development of the massive parallel sequencing technologies. Now we can obtain the complete genetic information of a person and analyse how this information is being used in a few days and with a limited cost. This technology makes possible a kind of research that was unthinkable a few years back. Our group, combining experimental and computational work, aims to develop and apply novel genome-wide techniques to address fundamental biological questions with medical implications. We are especially interested in how subtle variations in gene expression can lead to differential cellular phenotypes in humans and microorganism. Specifically, we investigate:
- Molecular bases of non-genetic cellular heterogeneity. We focus on understanding how single-cell variability, cellular plasticity and transcriptional memory contribute to the appearance of drug-tolerant cancer persister cells (i.e. those cells that, although genetically sensitive to a drug, do not respond to it). To reach that goal we combine the dissection of genetic factors controlling non-genetic heterogeneity with the development of novel genome-wide technologies to study this process.
- Crosstalk between ribosome dynamics and mRNA degradation. We have previously shown how the existence of widespread co-translational mRNA degradation allows to study ribosome dynamics by sequencing mRNA degradation intermediates (5P-Seq). Using that work as staring point, we to dissect the molecular crosstalk between mRNA degradation and ribosome dynamics in multiple organisms. We are characterizing how alterations in the translation process modulates mRNA stability and explore the utility of mRNA degradation signatures as reporters for cellular fitness in multiple organisms (from bacteria to cancer cells).
- Novel tools for molecular diagnosis. We are using our genomics expertise to improve and develop new molecular diagnosis and clinical genomic tools. We develop sequencing-based approaches to improve cancer-patient stratification and to accelerate the diagnosis of antimicrobial resistant infections.
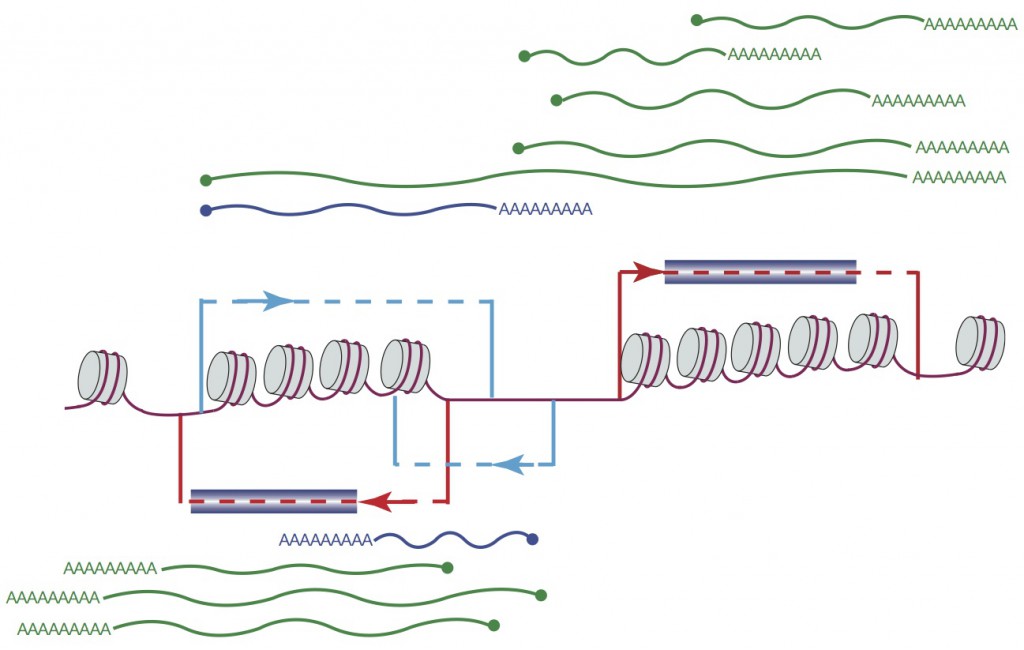
Figure 1: Extensive variation in transcript start and end sites revealed by TIF-Seq, a novel technique for transcript isoform profiling (Pelechano et al., Nature 2013)
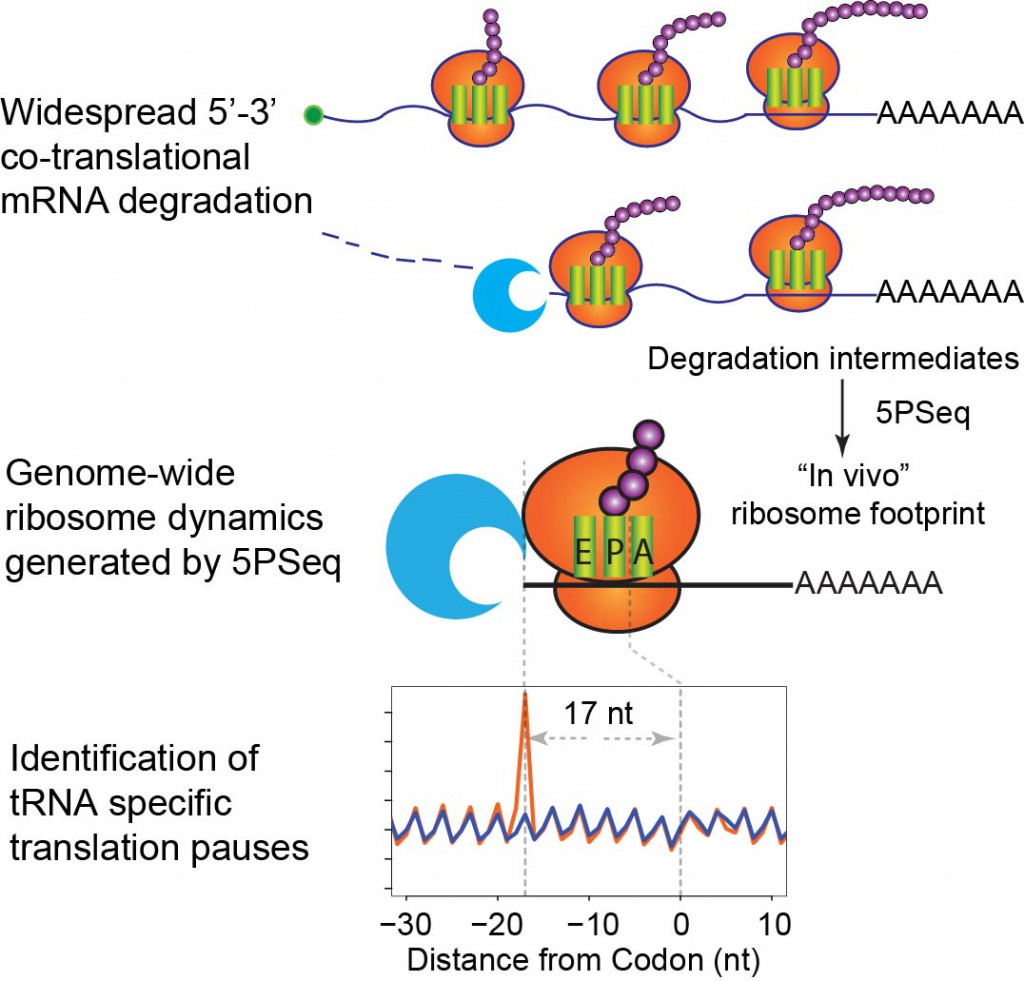
Figure 2: Widespread co-translational mRNA decay produces an in vivo ribosomal footprint and reveals ribosome dynamics (Pelechano et al. 2015 Cell)
Introduction to our research
Research highlight
We are honoured that the Knut and Alice Wallenberg Foundation drafted a research highlight about our work on June 2022. Thanks to the support from the Wallenberg foundation, our group can focus on long-term high-impact research.
Differences between identical cells offer potential for better cancer therapies
Despite possessing identical genes, cells can react differently when exposed to the same stimuli. This phenomenon can enable cancer cells to survive treatment that should kill them. Vicent Pelechano is studying the mechanisms involved to obtain knowledge that may lead to better cancer therapies.
Photo Magnus Bergström. Text Sara Nilsson.

